Biologists use genetic analysis to pinpoint source of cheese contamination

By carefully analyzing the genetics of specific bacteria killers, a BYU research team has been able to pinpoint where cheese contamination starts.
A new study led by BYU biology professor Keith Crandall details this method of genetic analysis as used to identify the source of cheese contamination in dairy factories. The study will appear in a forthcoming issue of Genome Biology and Evolution.
“The data are specific to these factories, but the techniques we’re developing are applicable anywhere,” Crandall said. “This kind of contamination happens all over the place and if you want to do reasonable intervention, you need to know the avenues of contamination.”
Generally speaking, bacteria are bad – unless you’re trying to make cheese.
In that case, bacteria serve as key agents in turning milk into cheddar, mozzarella or Gouda. So when something is killing off helpful dairy bacteria, there can be costly delays in the cheese-making process.
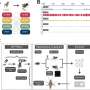
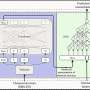
Carrying out a host of analyses, Crandall’s team traced the migration routes of the virus-like bacteria killers and found the contamination was being spread by humans; it wasn’t a case of the cheese-killer itself multiplying and spreading.
“There was one particular milk supplier who was contaminated, and using these approaches we could identify who that was,” Crandall said. “In the end you go back and disrupt the transmission patterns to make sure you don’t get contamination. It’s much easier to stop contamination once you figure out how it’s happening.”
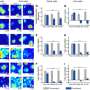
Crandall and his team analyzed 28 complete genomes of the bacteria killers, which are known as bacteriophage, or phage for short. These phage were sampled from eight Australian dairy factories over an 8-year period.
In addition to tracking the migration patterns, Crandall’s team dissected the genome data to see how the phage was adapting to attack the good bacteria. Phage are particularly troublesome entities because they evolve so quickly and often lead the arms race in the battle to eradicate them.
“But you can basically grow bacteria in the presence of these kinds of phage with these mutations and evolved resistance in the bacteria,” Crandall says. “It turns out that bacteria evolve pretty fast too.”
In cheese factories, phage-induced delays to fermentation can cause significant difficulties in a process that is based on milk, a perishable material that can’t be stored in the event of delay. Phage infections can also lead to negative repercussions in flavor and texture and can result in significant economic losses.
The factories involved in the study use anywhere from 100,000 liters to more than 1 million liters of milk a day. In other words, delays in the process means a lot of wasted milk.
The cheese-dairy-bacteriophage issue is nothing new. Rodney Brown, dean of the BYU College of Life Sciences, published a paper on the subject back in 1984.
“This has been a problem for a very long time,” Brown said. “But we hardly understood the problem like we do now. Now we have the tools and techniques to solve these kinds of problems that were only dreamed of back then.”
Crandall has used the same genetic analysis he used on the bacteriophage to analyze HIV, H1N1, Rhinovirus and others.
“You use these exact same techniques when you have an outbreak of whatever the next disease is,” he said.
BYU graduate student Eduardo Castro-Nallar was the lead author of the study.
Provided by Brigham Young University