Cancer stem cell marker also drives transcription in normal cells
New research links the recently discovered function of a multi-faceted transcriptional complex to control of gene expression in both normal cells and cancer stem cells. Two separate studies, published by Cell Press in the January 18th issue of Molecular Cell, provide insight into novel subunits associated with an evolutionarily conserved transcriptional regulatory complex and reveal a previously undescribed chromatin function that is required for full activity of nuclear receptors in normal cells and for the MYC oncoprotein in tumor cells.
Initiation of transcription requires sophisticated coordination of many different regulatory factors. Coactivators are multi-subunit complexes that facilitate transcription initiation directly, by interacting with RNA polymerase and general transcription factors, or indirectly, by influencing chromatin. For example, histone acetyltransferase (HAT) complexes are thought to activate gene expression by modifying chromatin-associated proteins called histones which function like spools for DNA to wind around.
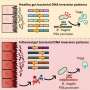
The yeast SAGA complex and the homologue metazoan TFTC/STAGA, also called hSAGA, are HAT-containing complexes that facilitate access of general transcriptional factors to DNA through histone acetylation. Although hSAGA is thought to be a homologue of the well-studied yeast SAGA complex, its subunit composition and functions are not as well understood. Dr. Didier Devys from the Institute de Génétique et de Biologie Moléculaire et Cellulaire in Strasbourg, France and colleagues identified three novel subunits, ATXN7L3, USP22 and ENY2, that are homologues of previously described subunits in the yeast SAGA complex.
The researchers demonstrated that the newly identified subunits work together to remove the ubiquitin moiety from monoubiquitylated histone H2B, similarly to what has been previously described in yeast, but also remove the ubiquitin moiety from monoubiquitylated histone H2A. The latter modification is not found in yeast but is more prevalent than monoubiquitylated H2B in mammals. Importantly, the deubiquitylation module of the Drosophila TFTC/STAGA complex was an enhancer of position effect variegation and counteracted heterochromatin silencing while both the Drosophila and the human deubiquitylation module were shown to be required for full transcriptional activation by the androgen receptor. This finding is clinically significant as androgen receptor activity is often deregulated in prostate cancer.
“The association of both HAT and deubiquitylation activities in the hSAGA complex provide an attractive mechanism by which the so called “cross-talk” between given histone marks is coordinated within the same regulatory complex” says Dr. Devys. “Further mechanistic studies are essential to examine the exact link between these activities and other chromatin modifying complexes to understand how these sequential events participate in chromatin remodeling and gene activation.”
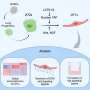
Working in parallel, a second research group led by Dr. Steven B. McMahon from Thomas Jefferson University’s Kimmel Cancer Center in Philadelphia also identified USP22 as a member of hSAGA. Previous work had identified USP22 as part of an eleven gene cancer stem cell signature that accurately distinguished patients whose tumors would eventually metastasize from those whose tumors would remain localized. “Unlike the other genes in this cancer stem cell signature, no direct mechanistic link to human cancer has been ascribed to USP22,” explains Dr. McMahon. McMahon’s group demonstrated that USP22 is required for activation of target gene transcription by the MYC oncoprotein and that USP22 depletion compromises MYC functions, including transformation of mammalian cells, and leads to cell cycle arrest.
Taken together, these results significantly advance the understanding of mechanisms that permit fine-tuning of transcriptional regulation by revealing that the hSAGA histone acetyltransferase complex is also capable of histone deubiquitylation. The findings provide critical new information about the importance of the timing and sequence of chromatin modifications in the control of gene expression in normal cells and shed light on the biochemical function of cancer stem cell marker and hSAGA subunit USP22, identifying it as a potential therapeutic target.
Source: Cell Press