Researchers develop database to help accelerate drug discovery
Researchers from Mount Sinai School of Medicine have developed a new computational method that will help streamline the analysis of gene expression experiments and provide scientists with a better mechanistic understanding of the differences between diseased and normal cells. The new database and software, called ChIP Enrichment Analysis (ChEA), will revolutionize how researchers identify drug targets and biomarkers.
Researchers can find the tool online at http://amp.pharm.mssm.edu/lib/chea.jsp. The data are published in the September 15th issue of Bioinformatics.
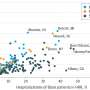
Until ChEA was developed, there was no centralized database that integrated results from ChIP-seq and ChIP-chip experiments—two types of experiments used to identify how proteins called transcription factors potentially regulate all genes in humans and mice. The data that results from these types of experiments is substantial and computational biologists have struggled with how to integrate them. Led by Avi Ma'ayan, PhD, Assistant Professor, Pharmacology and Systems Therapeutics, a team at Mount Sinai integrated the results from these experiments, collecting data from more than 100 proteins that bind DNA to regulate gene expression, called transcription factors, into one database.
"Using ChEA, we were able to link these transcription factors to the genes they regulate for the first time," said Dr. Ma'ayan. "Our program allows researchers to identify which proteins are likely responsible for genetic changes that may cause disease. Using our database, researchers will be able to better identify drug targets."
Dr. Ma'ayan's team tested ChEA in several case studies. One case study looked at two independent publications that found signature sets of genes that indicate whether a breast tumor was benign or malignant. Both studies came up with lists of genes that were biomarkers of each tumor type. However, the list uncovered in one study did not match the list in the other study; in fact, there was very little overlap in the biomarkers discovered in each study. However, when Dr. Ma'ayan entered the two lists into ChEA, he determined that the genes on both lists have the same regulatory protein, which determines the aberrant expression of these biomarker genes.
"Our reanalysis of these two publications verifies the usefulness of this software and database in gaining a better mechanistic understanding of the role transcription factors play in controlling gene expression, and subsequently, the development of disease," said Dr. Ma'ayan.